没养龙虾(OpenClaw),先养个马(Hermes)来做生物信息学
AI 发展日新月异,还没来得及养龙虾,马(Hermes)又来了。“弃龙虾(OpenClaw)、选爱马仕(Hermes)” ,似乎正在形成共识。真是应了那句话:只要学得慢,就不用学。
既然如此,那么我们就先不管龙虾,今天先来安装一个马试试。
安装
我们先看一下 Hermes 的 GitHub 主页:https://github.com/nousresearch/hermes-agent
今天(2026-05-12),hermes 在 GitHub 上有 14.5 万颗星,这对于才火起来 1 个月左右的项目来说,已经非常成功了。
进入文档页面:https://hermes-agent.nousresearch.com/
我们复制安装命令:
curl -fsSL https://hermes-agent.nousresearch.com/install.sh | bash运行安装——我这里是 Linux 系统。在运行这条命令之前,先要确保系统安装了Python 3.11。
配置
运行hermes setup
我们选择第一项:Quick setup。
设置大模型提供商
我们这里选择使用 DeepSeek。
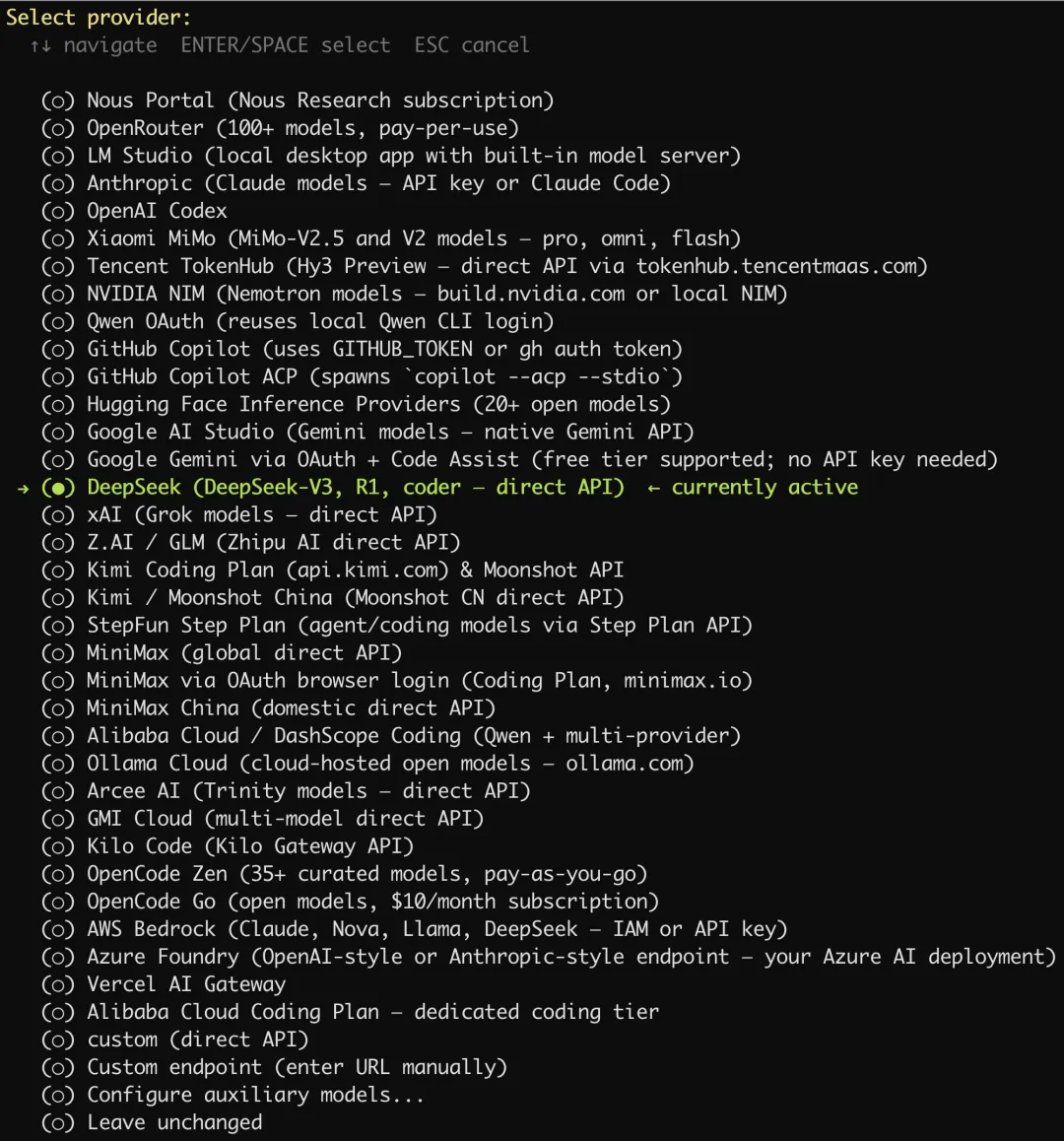
我们先到 DeepSeek 官网的开放平台申请好 API Key:

取一个名字,比如:hermes
然后回到终端,在这里输入刚才创建的 API key:
接着 Base URL 填写这个:https://api.deepseek.com
选择模型:
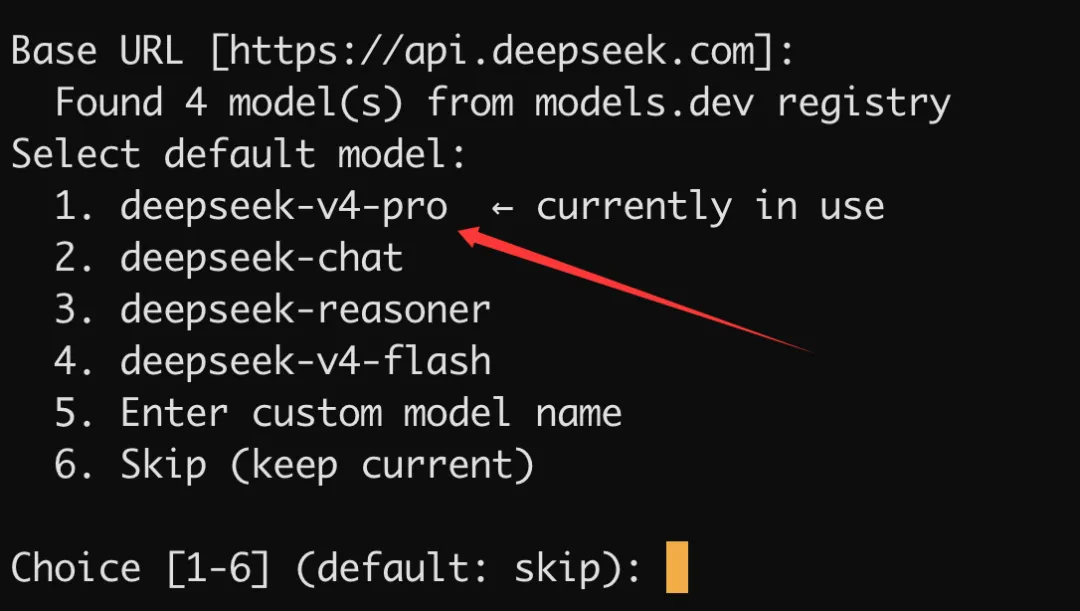
选择终端后端
保持默认选项就好了。
设置消息平台
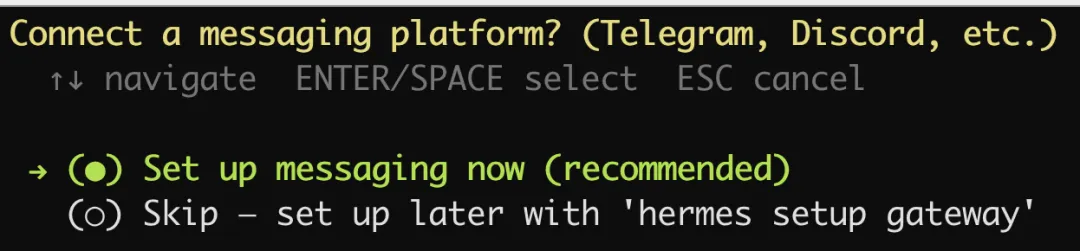
选择 QQ:
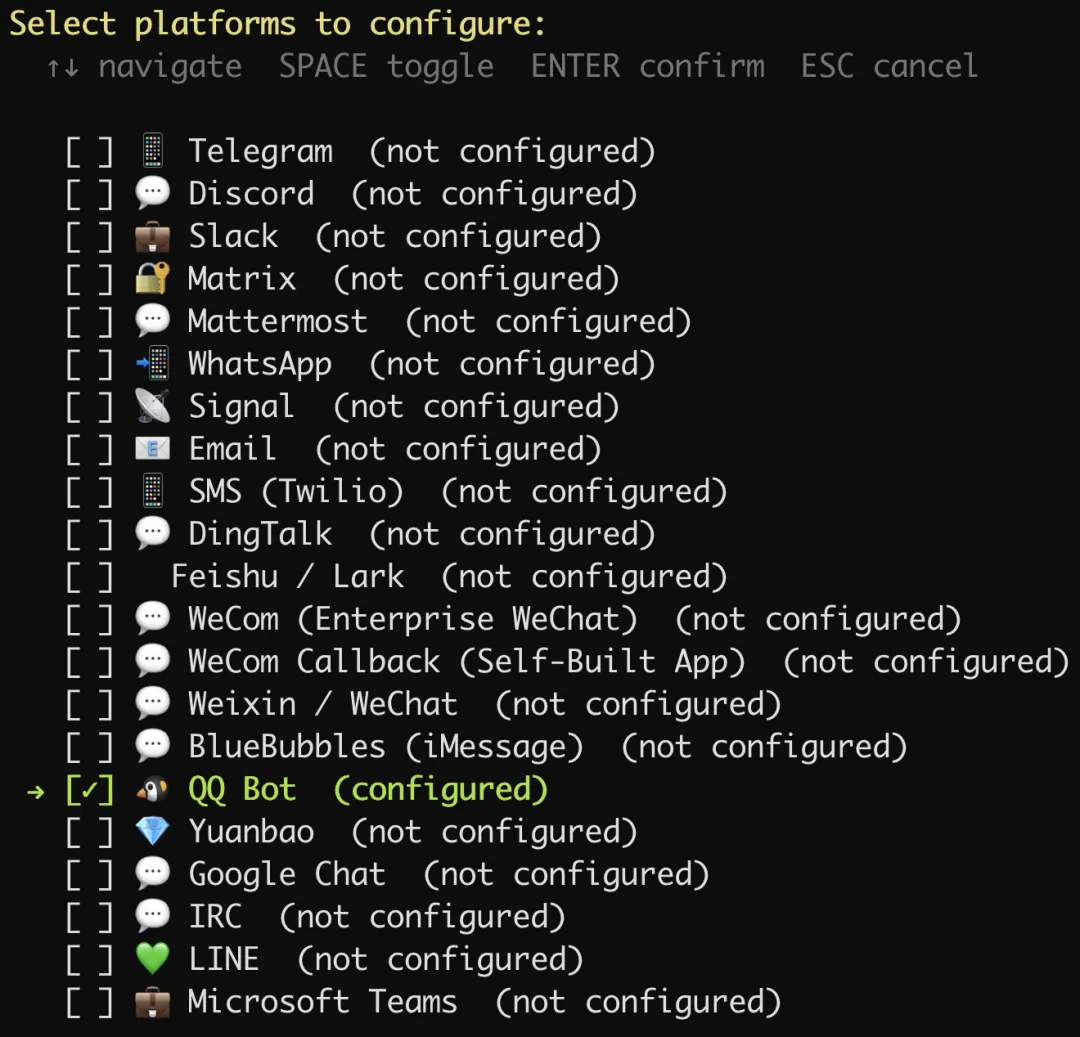
如果终端没有出现二维码,那么就复制提示的链接,到 QQ 开放平台创建一个 QQ 机器人,我们给它取一个名字:Hermes
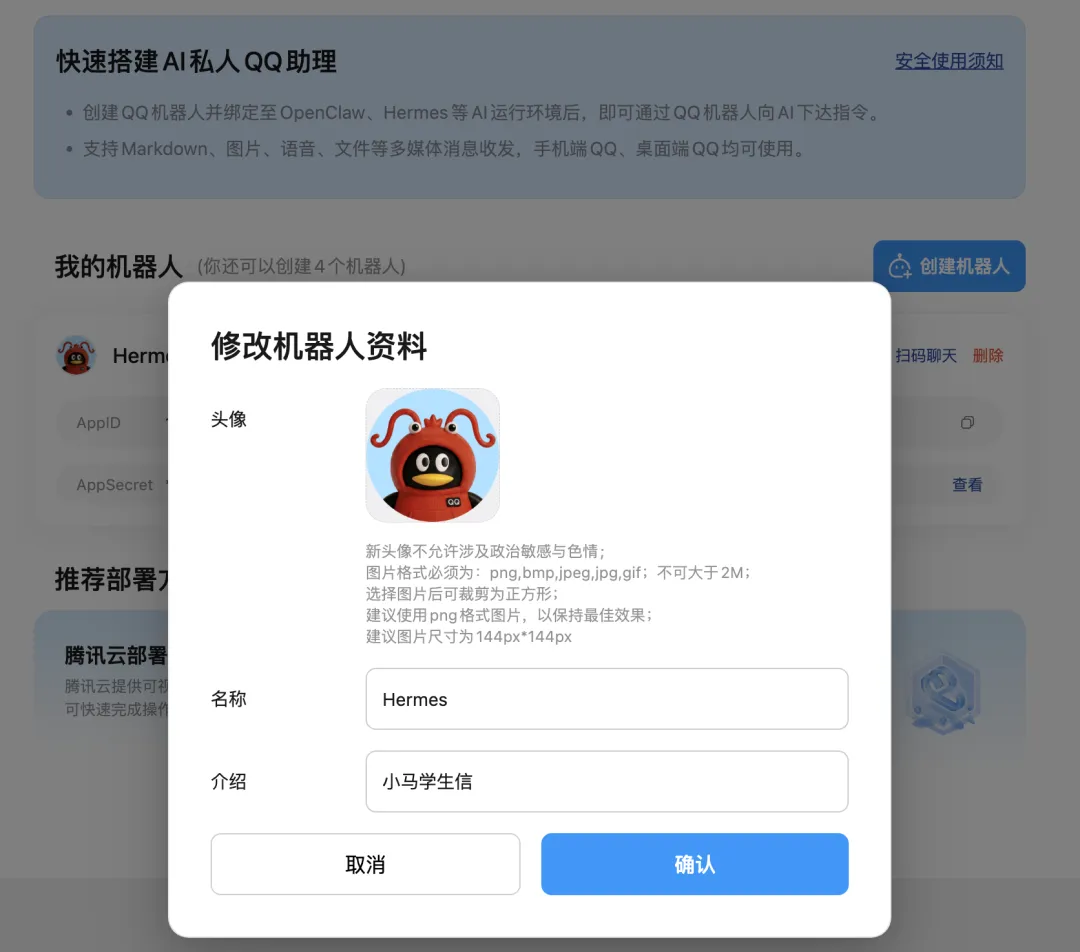
配置好之后:
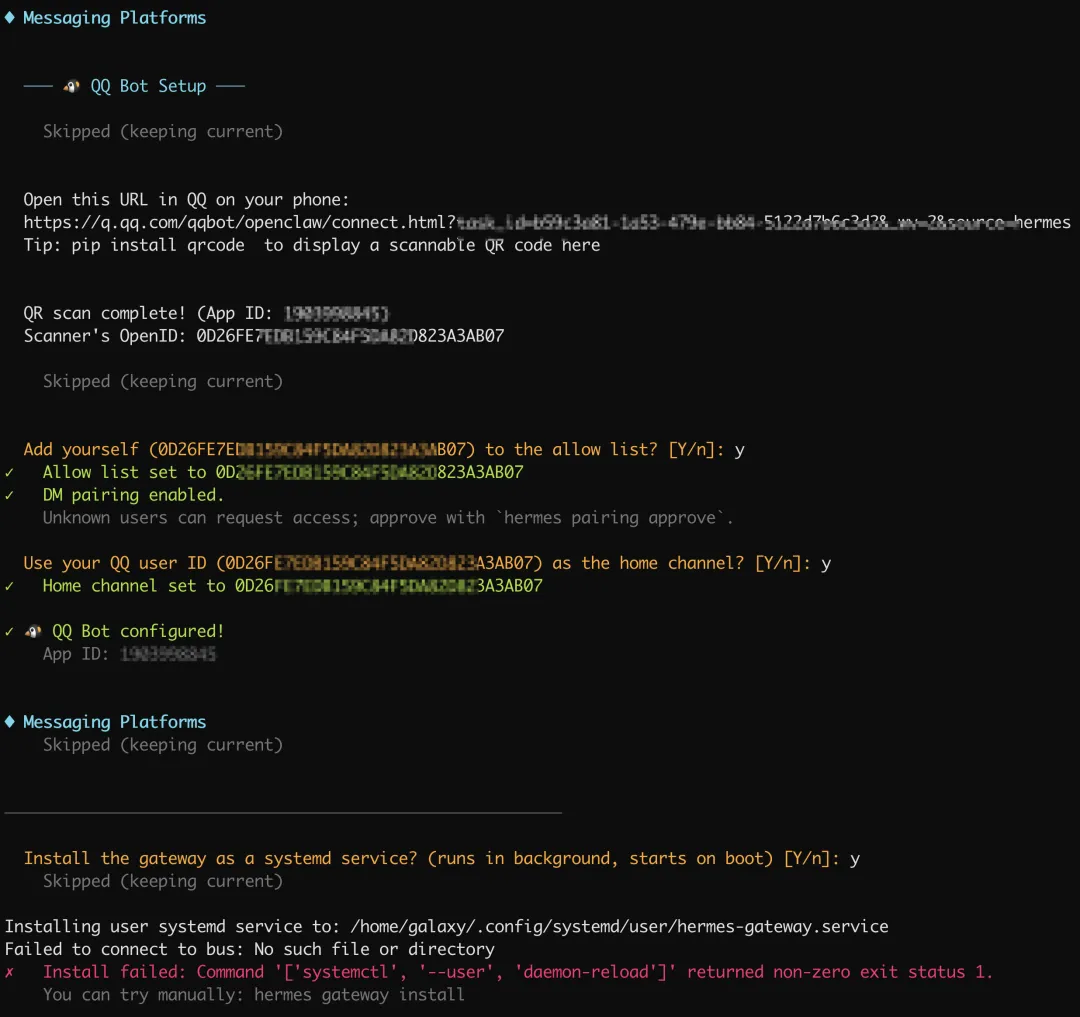
最后我们希望小马能常驻服务器,于是同意安装程序将其设为系统服务,这样重启服务器 hermes 也自动启动了。
但是出现了一条错误,原始是我们的当前账号galaxy没有添加系统服务的权限。
在 root 下执行:
# 1. 赋予 galaxy 用户常驻权限(确保退出登录后服务依然运行)loginctl enable-linger galaxy# 2. 切换到 galaxy 用户并进入对应目录su - galaxycd /home/galaxy/.hermes/hermes-agent# 3. 运行安装命令hermes gateway install打声招呼
要在 QQ 上跟 hermes 对话,首先必须要创建好机器人并且跟 hermes 配对好,就像蓝牙配对一样。其次hermes它只是一个智能体,它能工作要依赖于大模型,因此必须为其配置好大模型,我们这里用的是 DeepSeek v4。
可以看到,前面跟机器人对话它没反应,就是没有配置好。

单细胞学习路线图
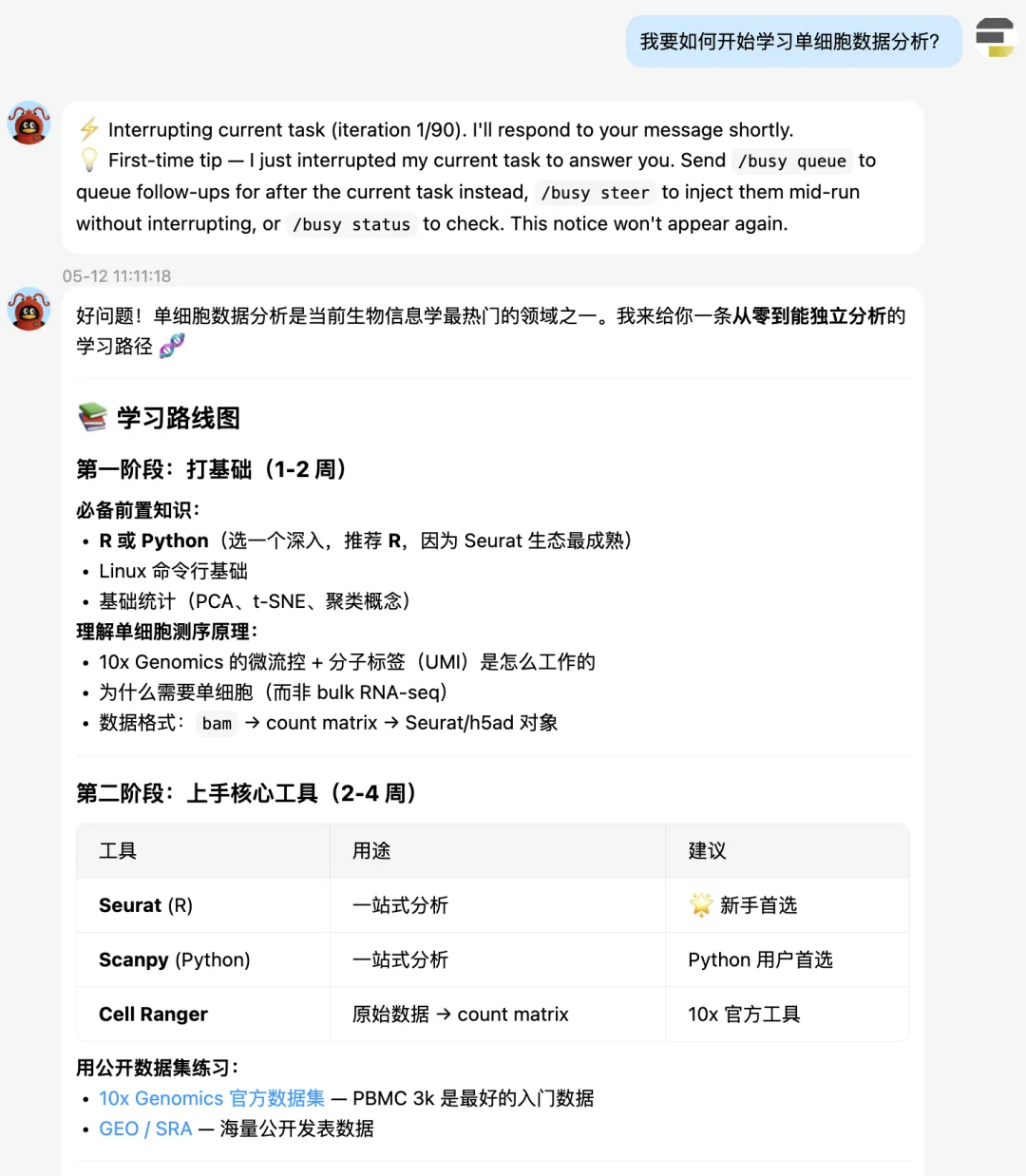


不得不说,hermes给的单细胞学习建议还是非常科学客观的。先是打语言基础,然后是核心工具的掌握:Seurat,Scanpy。接着是标准分析,进阶分析。
单细胞标准分析
大家接下来看hermes的实战表演吧。
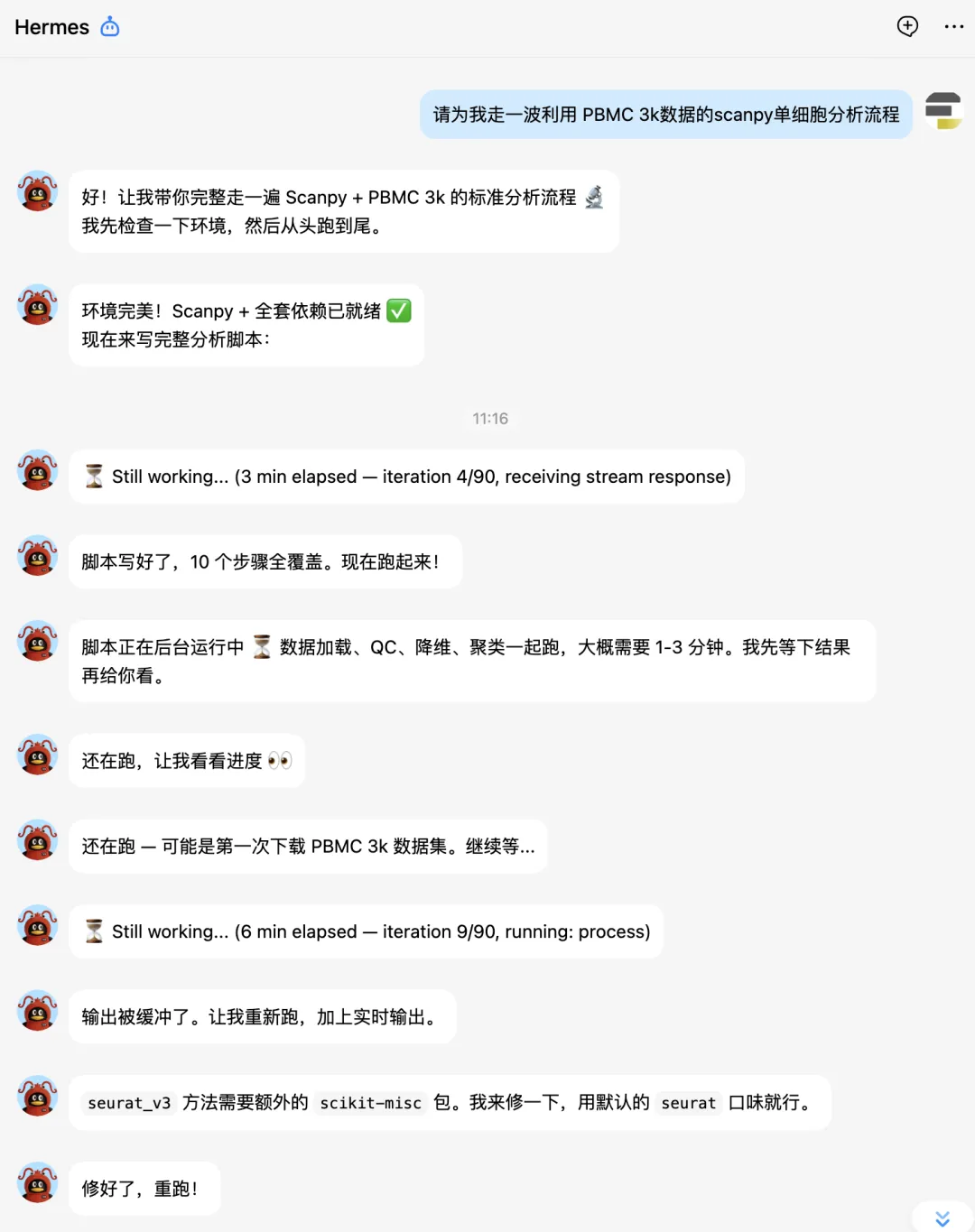
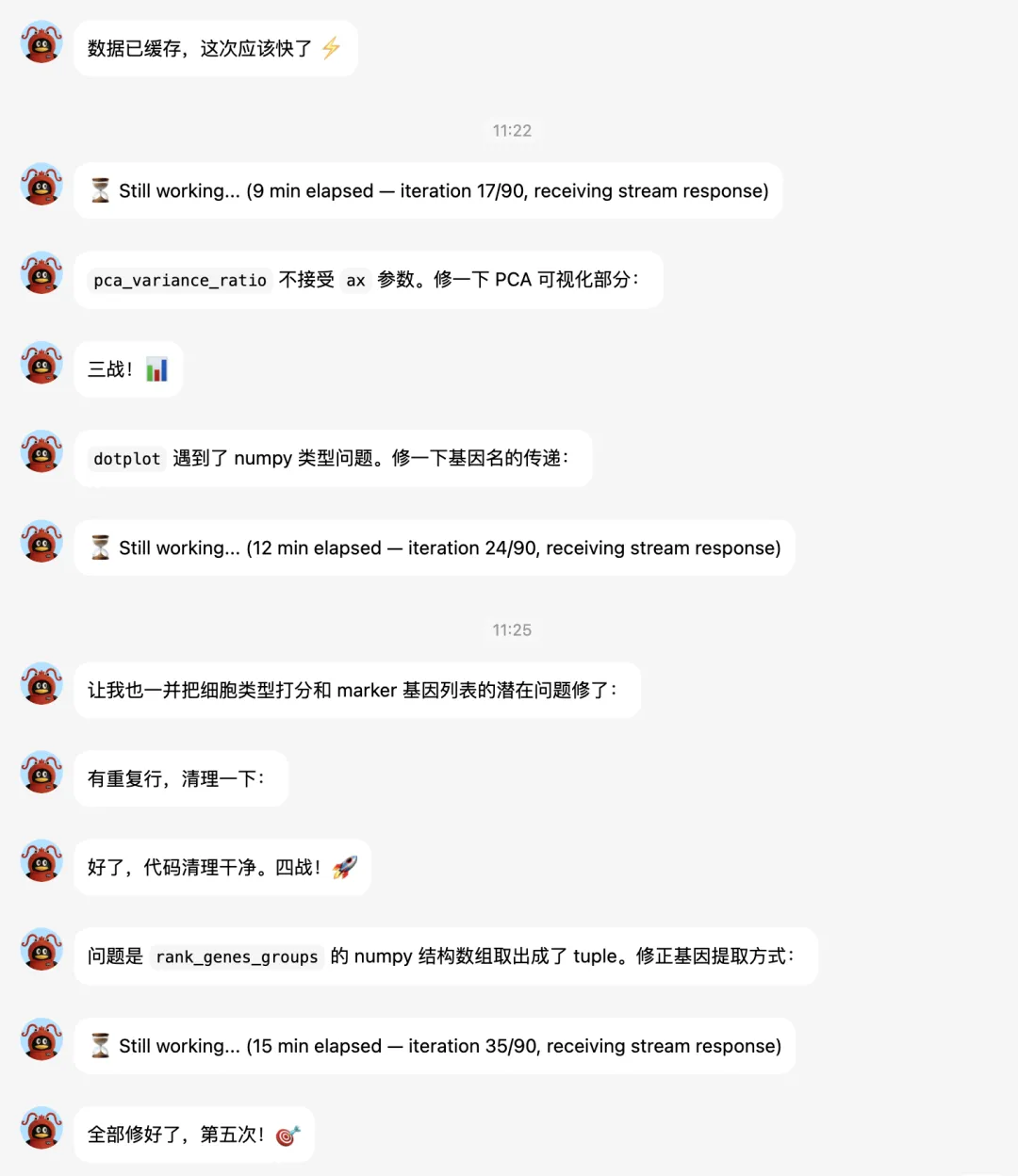
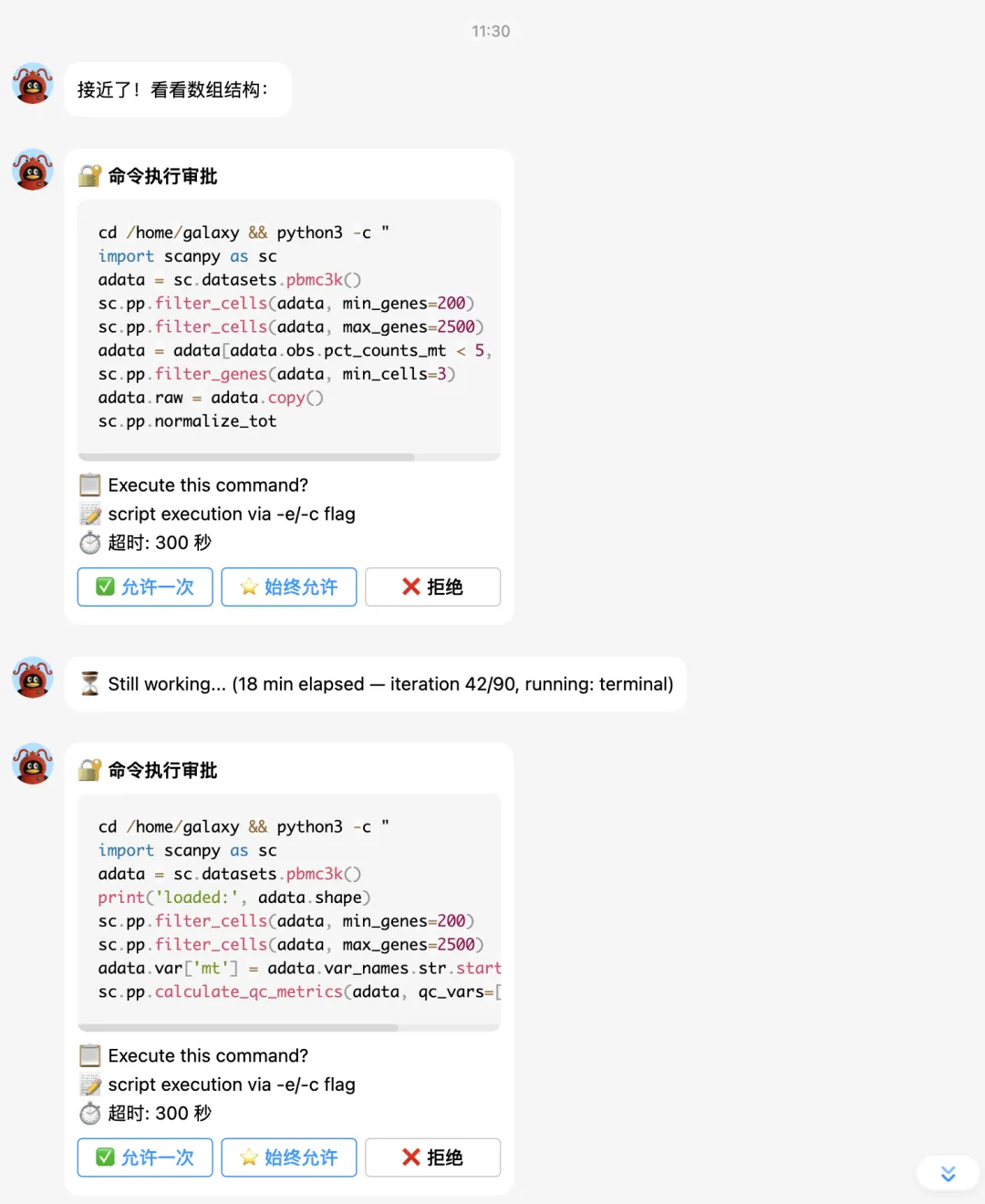
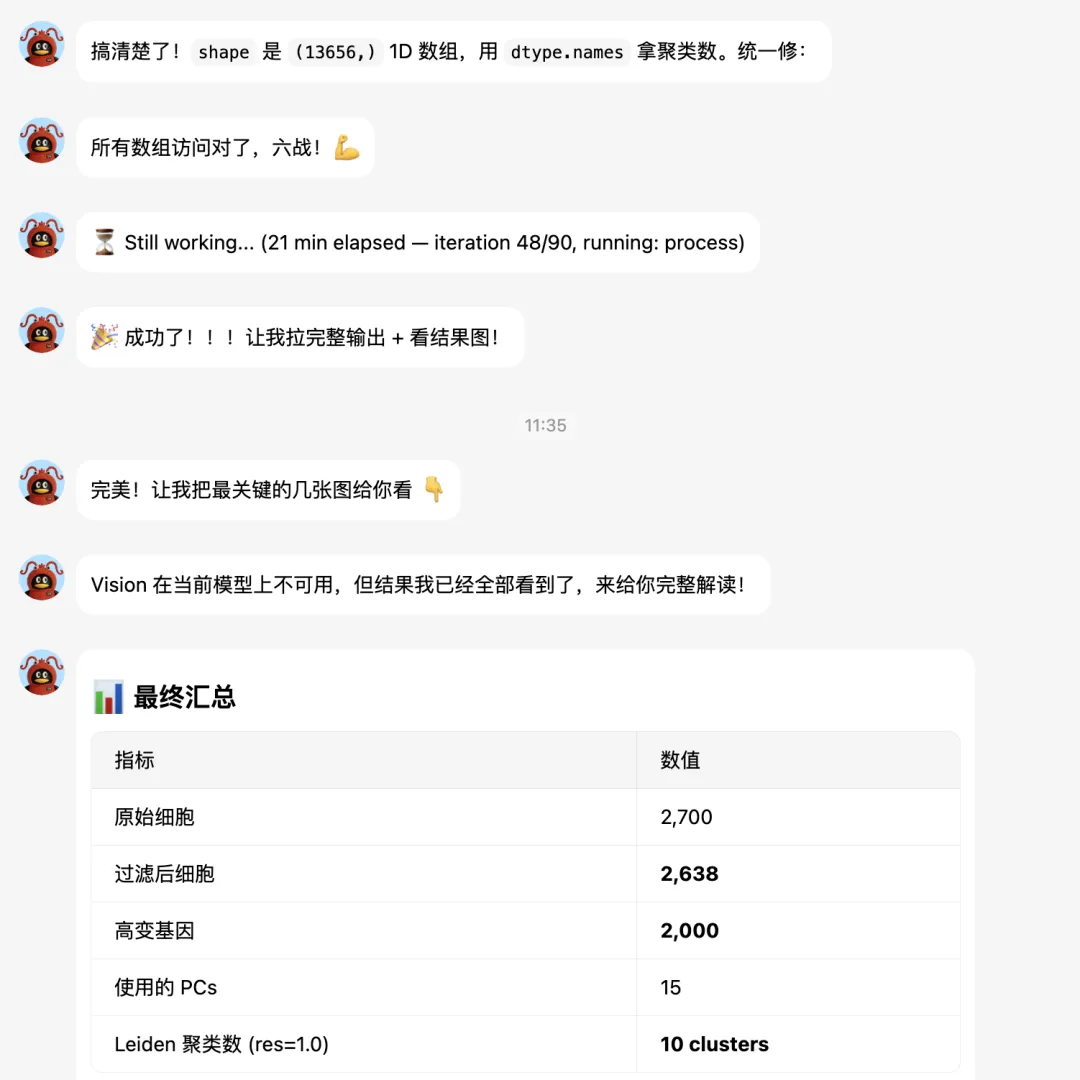
下面让它补一个celltypist分析:
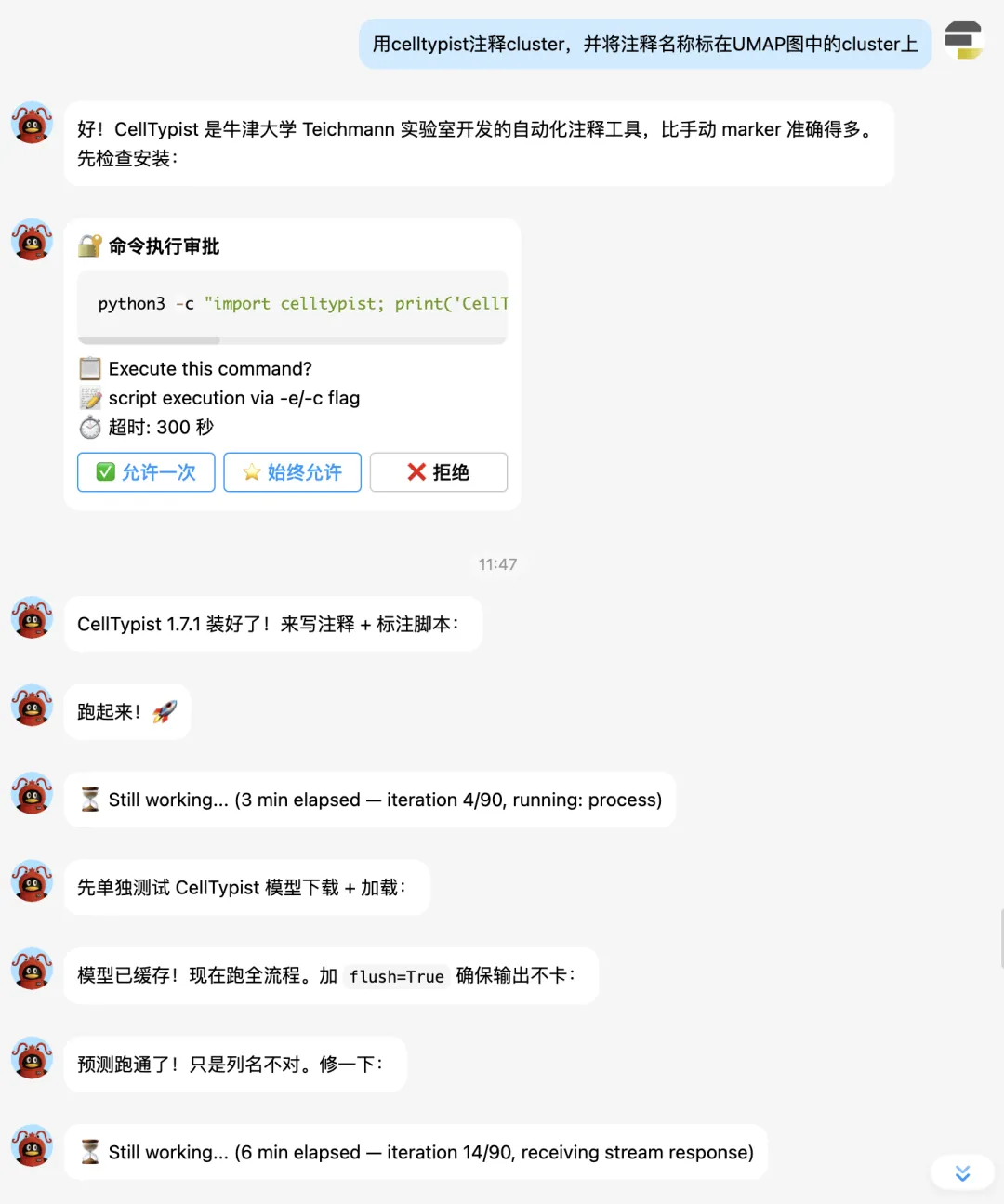
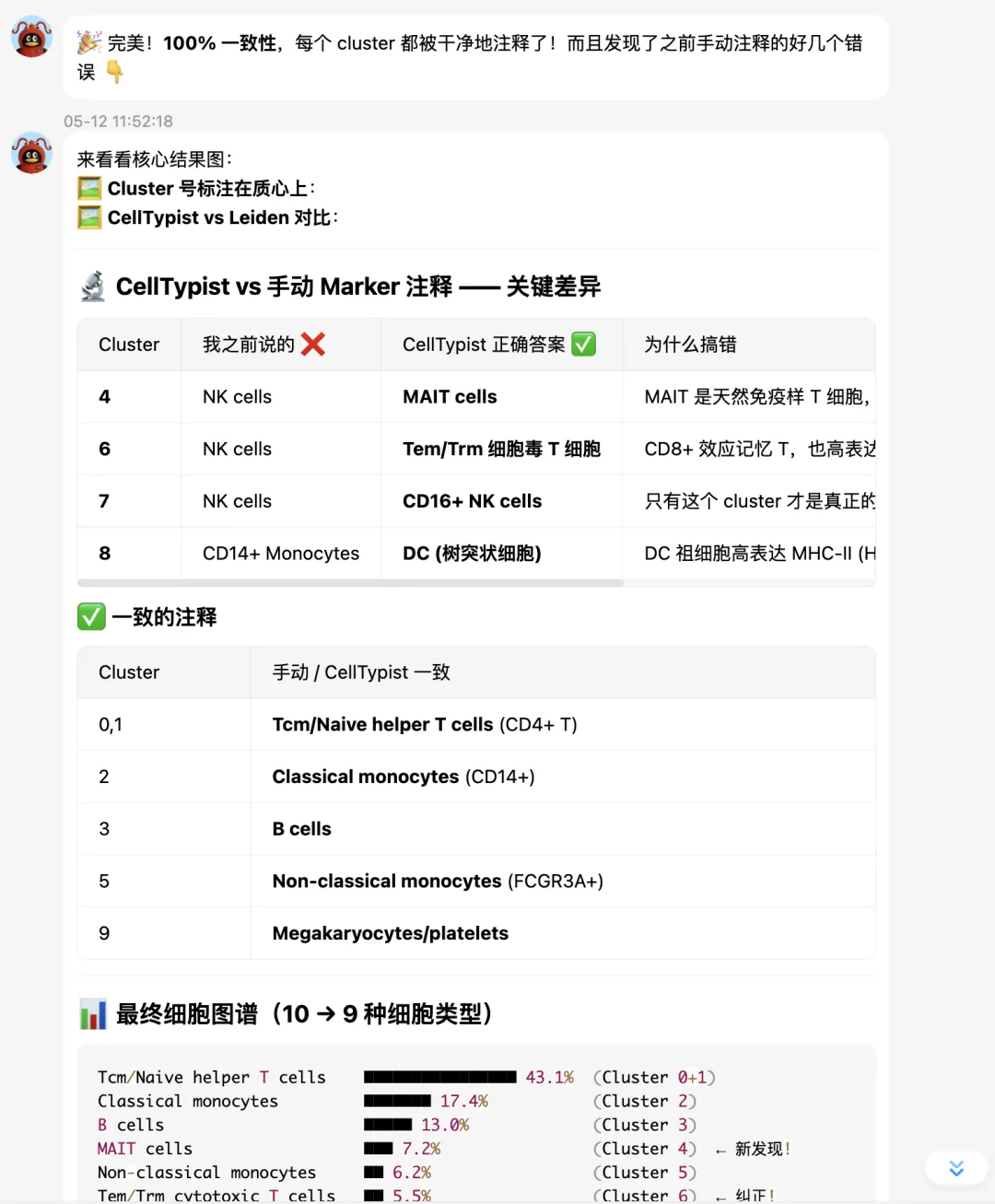
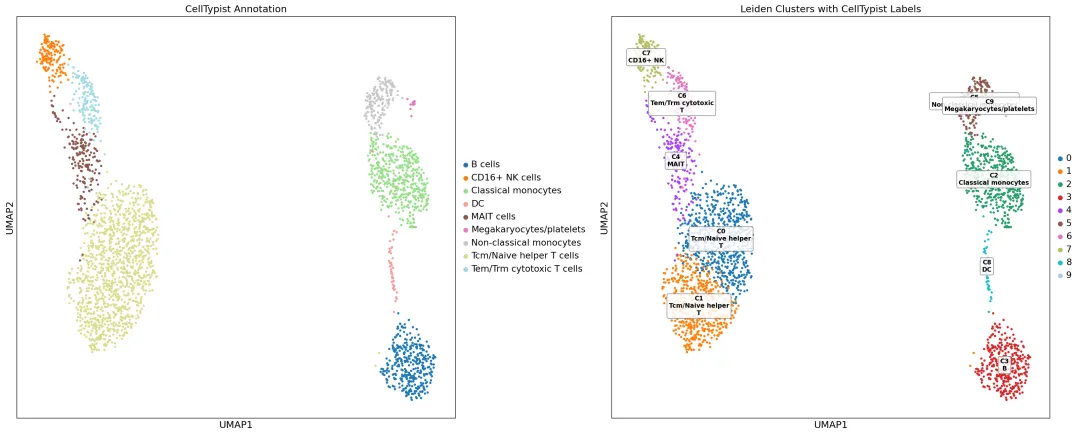
最终 UMAP 图:
最后总结
有意思的是,hermes会自动将分析过程提取成skill,这或许就是它跟OpenClaw很大的不同之处。后者更依赖人工编写skill。
比如我们刚才的分析就自动保存在
/home/galaxy/.hermes/skills/data-science/single-cell-scanpy目录下:
.├── references│ ├── celltypist_pitfalls.md│ └── rank_genes_groups_structured_array.md├── SKILL.md└── templates ├── celltypist_annotation.py └── pbmc3k_pipeline.py2 directories, 5 filesSKILL.md
---name: single-cell-scanpydescription: Single-cell RNA-seq analysis with Scanpy — QC, normalization, HVG, PCA, UMAP, clustering, marker detection, and cell-type annotation (manual marker + CellTypist automated).---# Single-Cell RNA-seq Analysis with ScanpyStandard scRNA-seq analysis pipeline using Scanpy (Python). Covers loading data through cell-type annotation, with tool-specific pitfalls documented below.## Triggers- User asks to run single-cell analysis, scRNA-seq, Scanpy, PBMC analysis- User mentions UMAP, Leiden clustering, marker genes, Seurat/Scanpy- User wants to analyze `.h5ad` files or 10x Genomics data## Standard Pipeline (10 Steps)1. **Load data** — `sc.datasets.pbmc3k()` or `sc.read_h5ad()`2. **QC filtering** — mark MT/ribo genes, `sc.pp.calculate_qc_metrics`, filter cells by n_genes and %MT3. **Normalization** — `sc.pp.normalize_total(target_sum=1e4)` + `sc.pp.log1p()`4. **HVG selection** — `sc.pp.highly_variable_genes(n_top_genes=2000)` (use default `seurat` flavor unless `scikit-misc` installed)5. **Regress + scale** — `sc.pp.regress_out(['total_counts', 'pct_counts_mt'])` then `sc.pp.scale(max_value=10)`6. **PCA** — `sc.tl.pca(svd_solver='arpack', n_comps=50)`7. **Neighbors + UMAP** — `sc.pp.neighbors(n_pcs=15)`, `sc.tl.umap()`8. **Clustering** — `sc.tl.leiden(resolution=1.0)`9. **Marker genes** — `sc.tl.rank_genes_groups(groupby, method='wilcoxon', use_raw=True)`10. **Cell-type annotation** — manual marker-based or automated via **CellTypist** (recommended: majority voting per cluster, far more accurate than manual markers)## CellTypist Automated Annotation (Step 10-b)CellTypist (Oxford Teichmann Lab) uses a pre-trained model with 98 immune cell types and 4164 gene features. **Always prefer this over manual marker-based annotation** — manual markers routinely misclassify MAIT cells, CD8+ Tem/Trm, and DCs (all of which express NKG7/CCL5/GZMB and get confused with NK cells).```pythonfrom celltypist import modelsmodel = models.Model.load(model='Immune_All_Low.pkl') # auto-downloads# CellTypist needs log-norm data; create clean copyadata_ct = adata.raw.to_adata()adata_ct.obs = adata.obs.copy()sc.pp.normalize_total(adata_ct, target_sum=1e4)sc.pp.log1p(adata_ct)predictions = celltypist.annotate( adata_ct, model='Immune_All_Low.pkl', majority_voting=True, over_clustering='leiden')adata.obs['celltypist_label'] = predictions.predicted_labels['majority_voting'].values```See `references/celltypist_pitfalls.md` for model output columns, data prep, and manual-vs-automated comparison.Full working script: `templates/celltypist_annotation.py`.## Critical Pitfalls### P1: `rank_genes_groups` yields structured numpy arrays- `adata.uns['rank_genes_groups']['names']` is a 1D structured recarray- Shape = `(n_genes,)` — NOT `(n_genes, n_clusters)`- Number of clusters = `len(names.dtype.names)` — NOT `shape[1]`- Access pattern: `names[rank][cluster_index]` — rank is gene rank (0=top), cluster_index is integer- Always cast genes to `str()` before passing to plotting functions```pythonnames = adata.uns['rank_genes_groups']['names']n_clusters = len(names.dtype.names)top_gene_cluster0 = str(names[0][0]) # top gene for cluster 0```### P2: `sc.pl.pca_variance_ratio` does not accept `ax` parameter- Use separate `plt.figure()` calls; save and close each individually- Same for `sc.pl.pca()` — it manages its own figure### P3: Dotplot `var_names` must be plain Python strings- Passing numpy record types (from structured arrays) causes `TypeError: unhashable type`- Always convert: `var_names=[str(g) for g in gene_list]`### P4: `sc.tl.score_genes` needs a flat gene list- Passing `list(marker_dict.values())` gives list-of-lists — wrong- Flatten: `[g for genes in marker_dict.values() for g in genes]`### P5: `sc.tl.rank_genes_groups` should use log-normalized data- Despite `use_raw=True`, log-normalize before calling or expect a warning- The pipeline above normalizes + log1p before HVG, so data is ready### P6: Vanilla `seurat` HVG flavor works out of the box- `flavor='seurat_v3'` requires `scikit-misc` (not commonly pre-installed)- Default `flavor='seurat'` needs no extra packages### P7: CellTypist — majority voting output has no `conf_score` column- `predictions.predicted_labels` columns: `['predicted_labels', 'over_clustering', 'majority_voting']`- Use `majority_voting` for cluster-level consensus; no separate confidence column- Must pass log-normalized data (not regressed/scaled) — create fresh `adata_ct` from `.raw`- Model auto-downloads on first use; wrap in `stdbuf -oL -eL` to avoid buffered-hang appearance## Running with real-time outputAlways use `stdbuf -oL -eL python3 -u script.py` or `PYTHONUNBUFFERED=1 python3 -u script.py` to avoid buffered stdout in long analyses.## Files| Path | Purpose || -------------------------------------------------- | ------------------------------------------------------------------ || `templates/pbmc3k_pipeline.py` | Complete 10-step pipeline — copy and modify for new datasets || `templates/celltypist_annotation.py` | CellTypist automated annotation with UMAP cluster labeling || `references/rank_genes_groups_structured_array.md` | Deep-dive on the structured array access pattern (hardest pitfall) || `references/celltypist_pitfalls.md` | CellTypist model output, data prep, manual-vs-automated comparison |pbmc3k_pipeline.py
分析代码保存成了模板:
#!/usr/bin/env python3"""PBMC 3k scRNA-seq Analysis Pipeline (Scanpy)==============================================Validated template for single-cell analysis. Replace dataset loadingin Step 1 to use your own .h5ad file.Generated by / updated by: see SKILL.md for full documentation."""import scanpy as scimport matplotlib.pyplot as pltimport os, sysimport warningswarnings.filterwarnings('ignore')sc.settings.verbosity = 2sc.settings.set_figure_params(dpi=100, facecolor='white', frameon=True)# --- Configuration ---OUT_DIR = "./results/"os.makedirs(OUT_DIR, exist_ok=True)# ============================================================# Step 1: Load data# ============================================================print("\n" + "="*60 + "\n Step 1: Load data\n" + "="*60)adata = sc.datasets.pbmc3k() # Replace with sc.read_h5ad('your_data.h5ad')print(f" Dimensions: {adata.shape[0]} cells x {adata.shape[1]} genes")# ============================================================# Step 2: Quality Control# ============================================================print("\n" + "="*60 + "\n Step 2: QC\n" + "="*60)# Mark mitochondrial and ribosomal genes (adjust prefix for your species)adata.var['mt'] = adata.var_names.str.startswith('MT-')adata.var['ribo'] = adata.var_names.str.startswith(('RPS', 'RPL'))sc.pp.calculate_qc_metrics(adata, qc_vars=['mt', 'ribo'], inplace=True)# --- QC plots (before filtering) ---fig, axes = plt.subplots(1, 3, figsize=(18, 5))sc.pl.violin(adata, ['n_genes_by_counts', 'total_counts', 'pct_counts_mt'], jitter=0.4, multi_panel=False, ax=axes[0], show=False)axes[0].set_title('QC (before)')sc.pl.scatter(adata, x='total_counts', y='n_genes_by_counts', color='pct_counts_mt', ax=axes[1], show=False)axes[1].set_title('Genes vs UMI')axes[2].hist(adata.obs['n_genes_by_counts'], bins=100, alpha=0.7)axes[2].axvline(200, color='red', linestyle='--', label='min=200')axes[2].axvline(2500, color='darkred', linestyle='--', label='max=2500')axes[2].set_xlabel('Number of genes')axes[2].set_title('Gene count distribution')axes[2].legend()plt.tight_layout()plt.savefig(f"{OUT_DIR}01_QC_before.png", dpi=150, bbox_inches='tight')plt.close()# --- Filtering ---print(f" Before filter: {adata.n_obs} cells")sc.pp.filter_cells(adata, min_genes=200)sc.pp.filter_cells(adata, max_genes=2500)adata = adata[adata.obs.pct_counts_mt < 5, :].copy()sc.pp.filter_genes(adata, min_cells=3)print(f" After filter: {adata.n_obs} cells, {adata.n_vars} genes")# QC after filteringfig, axes = plt.subplots(1, 3, figsize=(18, 5))sc.pl.violin(adata, ['n_genes_by_counts', 'total_counts', 'pct_counts_mt'], jitter=0.4, multi_panel=False, ax=axes[0], show=False)axes[0].set_title('QC (after)')sc.pl.scatter(adata, x='total_counts', y='n_genes_by_counts', color='pct_counts_mt', ax=axes[1], show=False)axes[1].set_title('Genes vs UMI (clean)')top20 = adata.var_names[adata.var['n_cells_by_counts'].argsort()[::-1][:20]]sc.pl.highest_expr_genes(adata, n_top=20, ax=axes[2], show=False)plt.tight_layout()plt.savefig(f"{OUT_DIR}02_QC_after.png", dpi=150, bbox_inches='tight')plt.close()# ============================================================# Step 3: Normalization# ============================================================print("\n" + "="*60 + "\n Step 3: Normalization\n" + "="*60)adata.raw = adata.copy()sc.pp.normalize_total(adata, target_sum=1e4)sc.pp.log1p(adata)# ============================================================# Step 4: Highly Variable Genes# ============================================================print("\n" + "="*60 + "\n Step 4: HVG selection\n" + "="*60)# Use default 'seurat' flavor (no extra deps). 'seurat_v3' needs scikit-misc.sc.pp.highly_variable_genes(adata, n_top_genes=2000, min_mean=0.0125, max_mean=3, min_disp=0.5)n_hvg = adata.var.highly_variable.sum()print(f" HVGs: {n_hvg}")sc.pl.highly_variable_genes(adata, show=False)plt.savefig(f"{OUT_DIR}03_HVG.png", dpi=150, bbox_inches='tight')plt.close()adata = adata[:, adata.var.highly_variable].copy()# ============================================================# Step 5: Regress + Scale# ============================================================print("\n" + "="*60 + "\n Step 5: Regress + Scale\n" + "="*60)sc.pp.regress_out(adata, ['total_counts', 'pct_counts_mt'])sc.pp.scale(adata, max_value=10)# ============================================================# Step 6: PCA# ============================================================print("\n" + "="*60 + "\n Step 6: PCA\n" + "="*60)sc.tl.pca(adata, svd_solver='arpack', n_comps=50)# PCA plots — note: pca_variance_ratio does NOT accept ax=sc.pl.pca_variance_ratio(adata, n_pcs=50, show=False)plt.title('Elbow plot')plt.savefig(f"{OUT_DIR}04a_elbow.png", dpi=150, bbox_inches='tight')plt.close()sc.pl.pca(adata, color=['n_genes_by_counts', 'pct_counts_mt'], show=False)plt.suptitle('PCA colored by QC metrics')plt.savefig(f"{OUT_DIR}04b_pca_qc.png", dpi=150, bbox_inches='tight')plt.close()# ============================================================# Step 7: Neighbors + UMAP# ============================================================print("\n" + "="*60 + "\n Step 7: Neighbors + UMAP\n" + "="*60)n_pcs = 15sc.pp.neighbors(adata, n_pcs=n_pcs, n_neighbors=15)sc.tl.umap(adata, min_dist=0.3, spread=1.0)# ============================================================# Step 8: Clustering# ============================================================print("\n" + "="*60 + "\n Step 8: Clustering\n" + "="*60)for res in [0.5, 0.8, 1.0, 1.2]: sc.tl.leiden(adata, resolution=res, key_added=f'leiden_r{res}')print(" Clusters per resolution:")for res in [0.5, 0.8, 1.0, 1.2]: print(f" res={res}: {adata.obs[f'leiden_r{res}'].nunique()}")adata.obs['leiden'] = adata.obs['leiden_r1.0'].astype(str)# UMAP clustering visualizationfig, axes = plt.subplots(2, 2, figsize=(14, 14))sc.pl.umap(adata, color='leiden', legend_loc='right margin', ax=axes[0,0], title=f'Leiden (n={adata.obs.leiden.nunique()})', show=False)sc.pl.umap(adata, color='n_genes_by_counts', ax=axes[0,1], title='n_genes', show=False)sc.pl.umap(adata, color='pct_counts_mt', ax=axes[1,0], title='%MT', show=False)sc.pl.umap(adata, color='total_counts', ax=axes[1,1], title='total UMI', show=False)plt.tight_layout()plt.savefig(f"{OUT_DIR}05_UMAP_clusters.png", dpi=150, bbox_inches='tight')plt.close()# ============================================================# Step 9: Marker Genes# ============================================================print("\n" + "="*60 + "\n Step 9: Marker Genes\n" + "="*60)sc.tl.rank_genes_groups(adata, 'leiden', method='wilcoxon', use_raw=True)# CRITICAL: rank_genes_groups['names'] is a 1D structured recarray# Shape = (n_genes,), clusters accessed via dtype.names# Access: names[rank][cluster_index]names_struct = adata.uns['rank_genes_groups']['names']scores_struct = adata.uns['rank_genes_groups']['scores']n_clusters = len(names_struct.dtype.names)print("\n Top5 markers per cluster:")for ci in range(n_clusters): clabel = names_struct.dtype.names[ci] markers = [str(names_struct[j][ci]) for j in range(min(5, len(names_struct)))] scores = [scores_struct[j][ci] for j in range(min(5, len(names_struct)))] print(f" Cluster {clabel:>2}: " + ', '.join(f"{g}({s:.1f})" for g,s in zip(markers, scores)))# Dotplot with top-3 per cluster (cast to plain strings!)top3_genes = []for ci in range(n_clusters): for rank in range(min(3, len(names_struct))): top3_genes.append(str(names_struct[rank][ci]))top3_genes = list(set(top3_genes))sc.pl.dotplot(adata, var_names=top3_genes, groupby='leiden', show=False)plt.savefig(f"{OUT_DIR}07_Dotplot_top3.png", dpi=150, bbox_inches='tight')plt.close()# ============================================================# Step 10: Cell-Type Annotation# ============================================================print("\n" + "="*60 + "\n Step 10: Cell-Type Annotation\n" + "="*60)# Adjust marker genes for your dataset/speciesmarker_dict = { 'CD14+ Monocytes': ['CD14', 'LYZ', 'S100A9'], 'FCGR3A+ Monocytes': ['FCGR3A', 'MS4A7', 'LST1'], 'CD4+ T cells': ['CD3D', 'CD3E', 'IL7R', 'CD4'], 'CD8+ T cells': ['CD3D', 'CD3E', 'CD8A', 'CD8B'], 'NK cells': ['NKG7', 'GNLY', 'KLRD1'], 'B cells': ['CD79A', 'MS4A1', 'CD19'], 'Dendritic cells': ['FCER1A', 'CST3'], 'Megakaryocytes': ['PPBP', 'PF4'],}# Average expression per cluster for each cell typecell_type_anno = {}for cluster in sorted(adata.obs['leiden'].unique(), key=int): mask = adata.obs['leiden'] == cluster ct_scores = {} for ct, genes in marker_dict.items(): valid_genes = [g for g in genes if g in adata.raw.var_names] if not valid_genes: ct_scores[ct] = 0 continue avg_expr = adata.raw[mask, valid_genes].X.mean() try: avg_expr = float(avg_expr) except: avg_expr = 0 ct_scores[ct] = avg_expr best_ct = max(ct_scores, key=ct_scores.get) cell_type_anno[cluster] = best_ct sorted_by_score = sorted(ct_scores.items(), key=lambda x: x[1], reverse=True) candidates = ' / '.join(f"{ct}({s:.2f})" for ct, s in sorted_by_score[:2]) print(f" Cluster {cluster:>2}: \u2192 {best_ct} ({candidates})")adata.obs['cell_type'] = adata.obs['leiden'].map(cell_type_anno)# Cell-type UMAPfig, axes = plt.subplots(1, 2, figsize=(20, 8))sc.pl.umap(adata, color='leiden', legend_loc='right margin', ax=axes[0], title='Leiden Clusters', show=False)sc.pl.umap(adata, color='cell_type', legend_loc='right margin', ax=axes[1], title='Predicted Cell Types', show=False)plt.tight_layout()plt.savefig(f"{OUT_DIR}09_UMAP_celltypes.png", dpi=150, bbox_inches='tight')plt.close()# Marker dotplot by cell typemarker_genes_all = list(dict.fromkeys( g for genes in marker_dict.values() for g in genes if g in adata.raw.var_names))sc.pl.dotplot(adata, var_names=marker_genes_all, groupby='cell_type', use_raw=True, dendrogram=True, show=False)plt.savefig(f"{OUT_DIR}10_Dotplot_markers.png", dpi=150, bbox_inches='tight')plt.close()# ============================================================# Summary# ============================================================print(f"""{'='*60} Analysis Complete!{'='*60}Cells after QC: {adata.n_obs}HVGs: {n_hvg}PCs used: {n_pcs}Leiden clusters: {adata.obs['leiden'].nunique()}Cell types:""")for ct in sorted(set(cell_type_anno.values())): count = (adata.obs['cell_type'] == ct).sum() pct = count / adata.n_obs * 100 print(f" {ct}: {count} cells ({pct:.1f}%)")print(f"\nResults saved to: {OUT_DIR}")大家可以看看它这个代码写得到底对不对,质量如何?
好了,今天养马的尝试就先到这里。大家有什么好的想法欢迎到群里交流:

中国银河生信云平台精品课程
单细胞数据分析培训班(Python/Galaxy可选),不怕学不会
转录组数据分析实战(Galaxy)| 直播回放 咨询小助手:usegalaxy
 夜雨聆风
夜雨聆风




